161762 Multivariate Analysis for Big Data
Lecture 6: Multidimensional Scaling and Factor Analysis
Massey University
Fall 2026
Learning objectives
Understand the basics of Multidimensional Scaling.
Understand the basics of Factor Analysis.
Refresher Distance Matrix
\[ \mathbf{X}_{n \times p} = \begin{bmatrix} x_{11} & \cdots & x_{1p} \\ \vdots & \ddots & \vdots \\ x_{n1} & \cdots & x_{np} \end{bmatrix} \Longrightarrow \mathbf{D} = \begin{bmatrix} 0 & d(x_1, x_2) & d(x_1, x_3) & \cdots & d(x_1, x_n) \\ d(x_2, x_1) & 0 & d(x_2, x_3) & \cdots & d(x_2, x_n) \\ d(x_3, x_1) & d(x_3, x_2) & 0 & \cdots & d(x_3, x_n) \\ \vdots & \vdots & \vdots & \ddots & \vdots \\ d(x_n, x_1) & d(x_n, x_2) & d(x_n, x_3) & \cdots & 0 \end{bmatrix} \]
Fill \(d(x_n,x_n)\) with your favorite distance method.
Choose your analysis: ordination or Clustering (later in the ocurse)
Multidimensional Scaling (MDS)
MDS & Related Methods - Historical Timeline
- 1952 | Torgerson introduces Metric MDS
- 1964 | Kruskal develops Non-metric MDS
- 1966 | Gower formalizes Classical MDS as Principal Coordinates Analysis (PCoA)
- 1971 | Gower publishes the Gower Similarity Coefficient (handles mixed data types)
Multidimensional Scaling (MDS)
Ordinate your samples in a reduce dimensional space.
MDS is an unconstrained ordination method.
Multidimensional Scaling (MDS)
| Athens | Cologne | Geneva | Hamburg | Istanbul | Lisbon | London | Madrid | |
|---|---|---|---|---|---|---|---|---|
| Athens | 0 | |||||||
| Cologne | 1212 | 0 | ||||||
| Geneva | 1068 | 323 | 0 | |||||
| Hamburg | 1268 | 227 | 540 | 0 | ||||
| Istanbul | 345 | 1239 | 1190 | 1236 | 0 | |||
| Lisbon | 1772 | 1149 | 929 | 1366 | 2003 | 0 | ||
| London | 1501 | 331 | 468 | 463 | 1562 | 972 | 0 | |
| Madrid | 1466 | 883 | 627 | 1107 | 1686 | 319 | 774 | 0 |
Multidimensional Scaling (MDS)
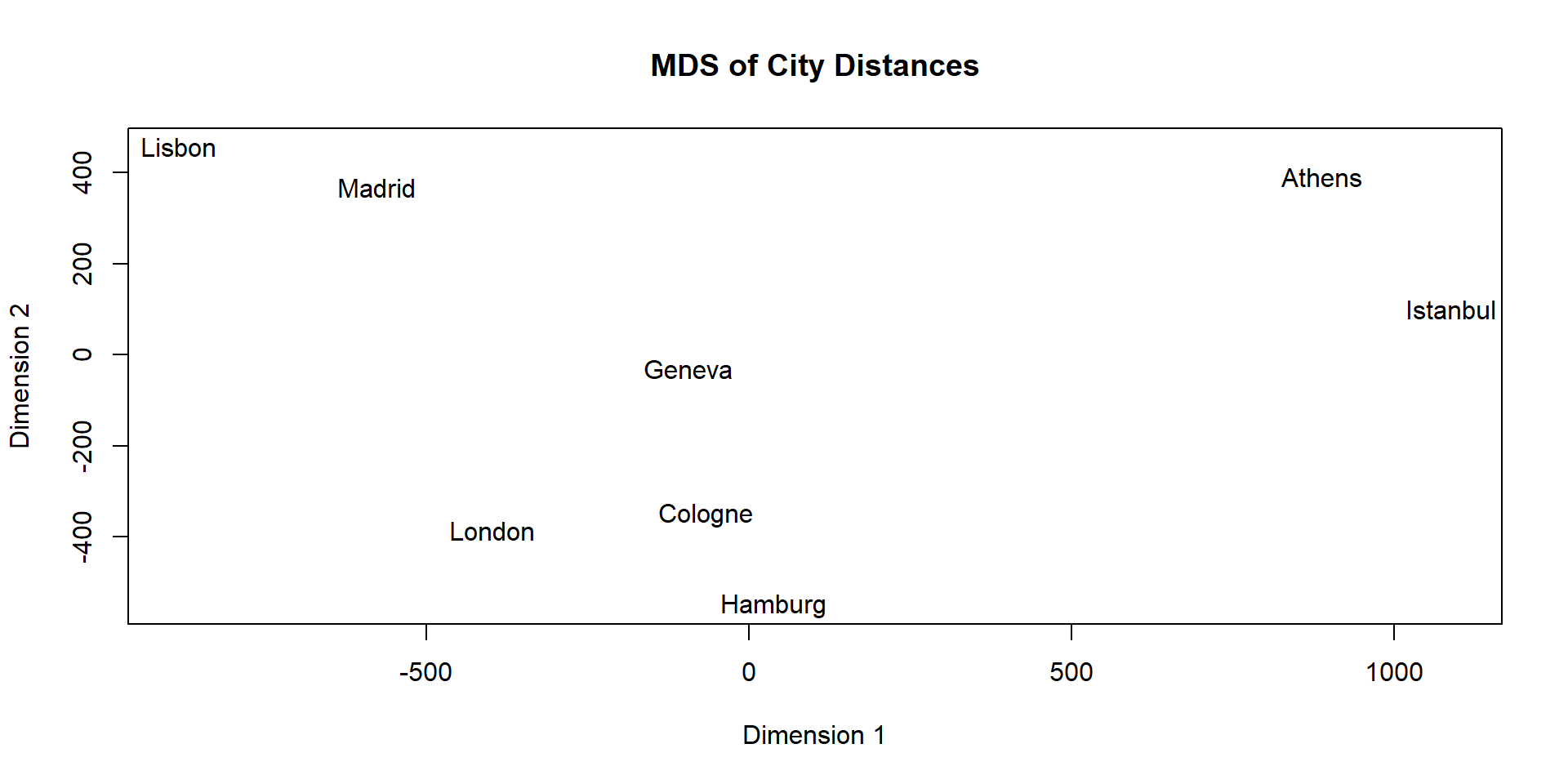

Multidimensional Scaling (MDS)
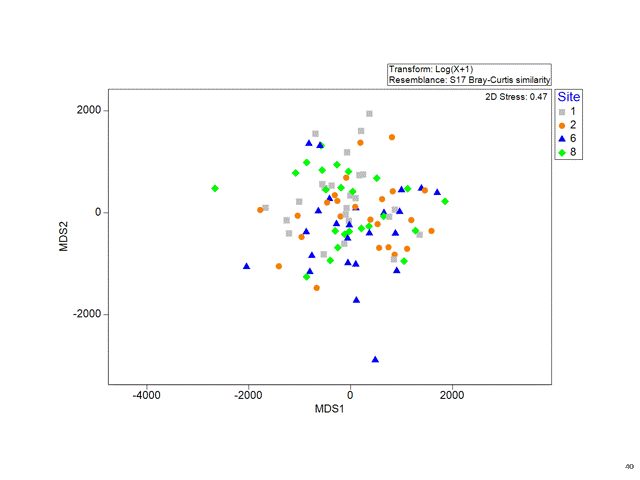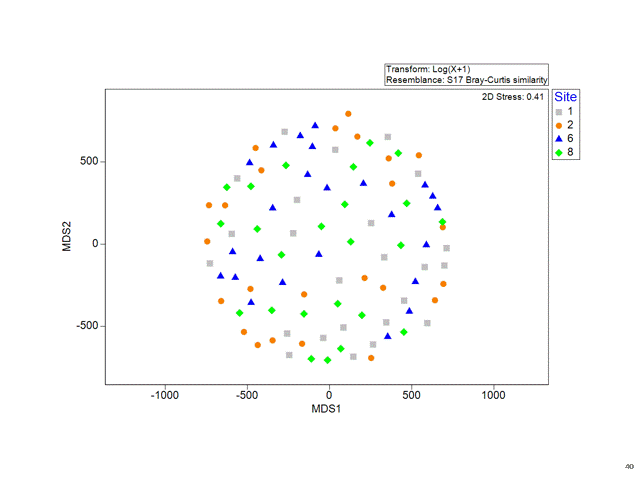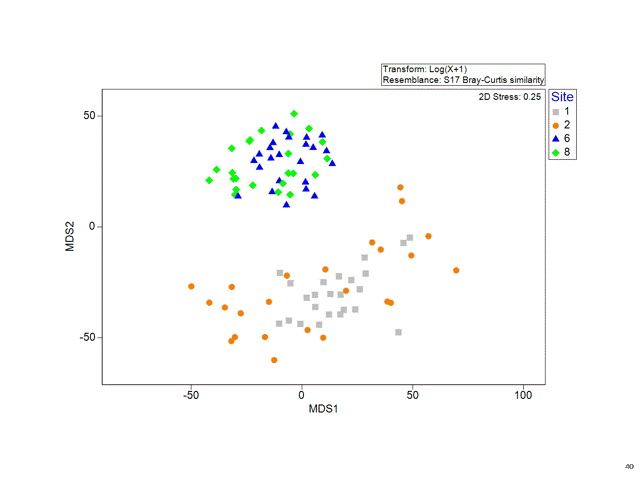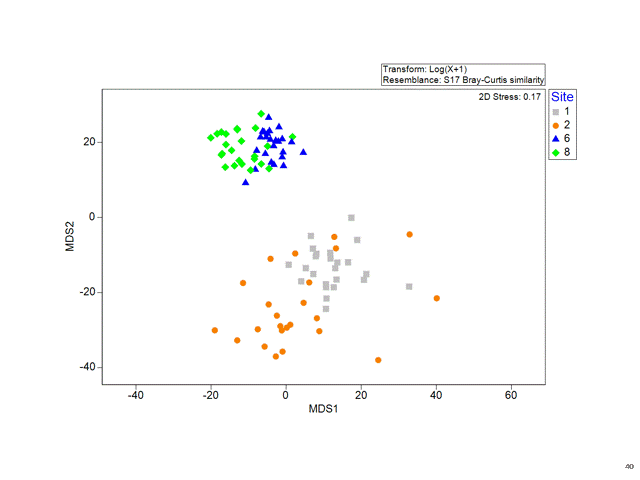
Multidimensional Scaling (MDS)
Original
| Ath | Col | Gen | Ham | Ist | Lis | Lon | Mad | |
|---|---|---|---|---|---|---|---|---|
| Ath | 0 | |||||||
| Col | 1212 | 0 | ||||||
| Gen | 1068 | 323 | 0 | |||||
| Ham | 1268 | 227 | 540 | 0 | ||||
| Ist | 345 | 1239 | 1190 | 1236 | 0 | |||
| Lis | 1772 | 1149 | 929 | 1366 | 2003 | 0 | ||
| Lon | 1501 | 331 | 468 | 463 | 1562 | 972 | 0 | |
| Mad | 1466 | 883 | 627 | 1107 | 1686 | 319 | 774 | 0 |
MDS
| Ath | Col | Gen | Ham | Ist | Lis | Lon | Mad | |
|---|---|---|---|---|---|---|---|---|
| Ath | 0 | |||||||
| Col | 1211 | 0 | ||||||
| Gen | 1069 | 324 | 0 | |||||
| Ham | 1269 | 225 | 538 | 0 | ||||
| Ist | 352 | 1241 | 1189 | 1236 | 0 | |||
| Lis | 1773 | 1149 | 927 | 1366 | 2003 | 0 | ||
| Lon | 1502 | 331 | 467 | 466 | 1563 | 973 | 0 | |
| Mad | 1466 | 883 | 626 | 1107 | 1687 | 318 | 775 | 0 |
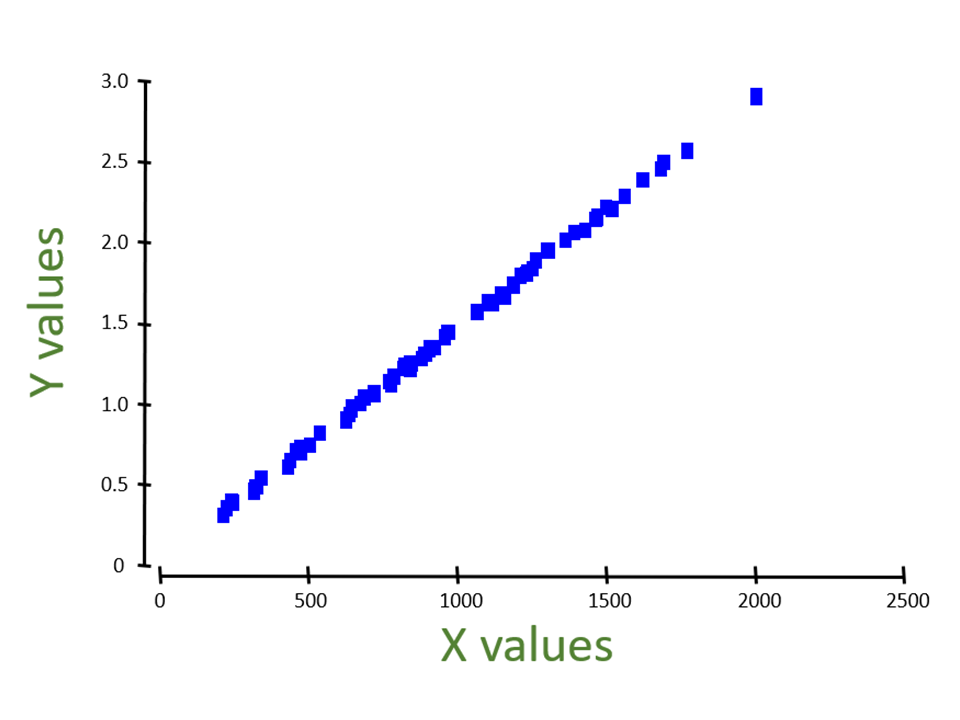
Adequacy of the ordination
PCA
- Euclidean distances in the ordination vs. Euclidean distances among the points based on the full set of (standardized) original variables
MDS
- Euclidean distances in the ordination vs. original dissimilarities or distances (based on a chosen measure).
Stress
\[ \text{S} = \frac{ \overbrace{\displaystyle \sum_{j<k} \left( d_{jk} - \delta_{jk} \right)^2}^{\text{Sum of squared residuals (MDS distance vs perfect regression)}} }{ \underbrace{\displaystyle \sum_{j<k} \delta_{jk}^2}_{\text{Sum of squared perfect regression distances (original distances)}} } \]
Stress
Is a measure of how good the fit is between the original distances and the new distances on the plot.
The lower the stress, the more the MDS plot reflects real distance structures in the data.
A published MDS plot should always state a value for stress. (if done iteratively).
Rule of thumb:
- Stress < 0.1 Good.
- 0.1 < Stress < 0.2 OK
- Stress > 0.2 Not so good
- Stress > 0.3 No better than random!
Metric VS Non-metric MDS (nMDS)
Metric MDS fits a strictly linear regression line (constant slope; fixed at the origin)
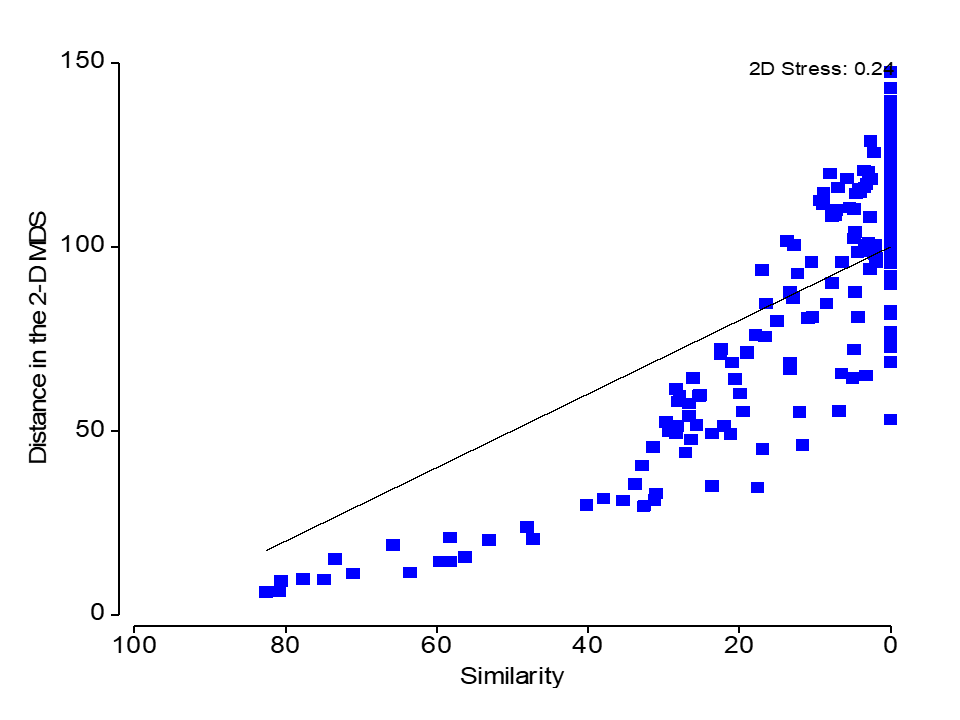
nMDS fits a monotonically increasing regression line (a step function)
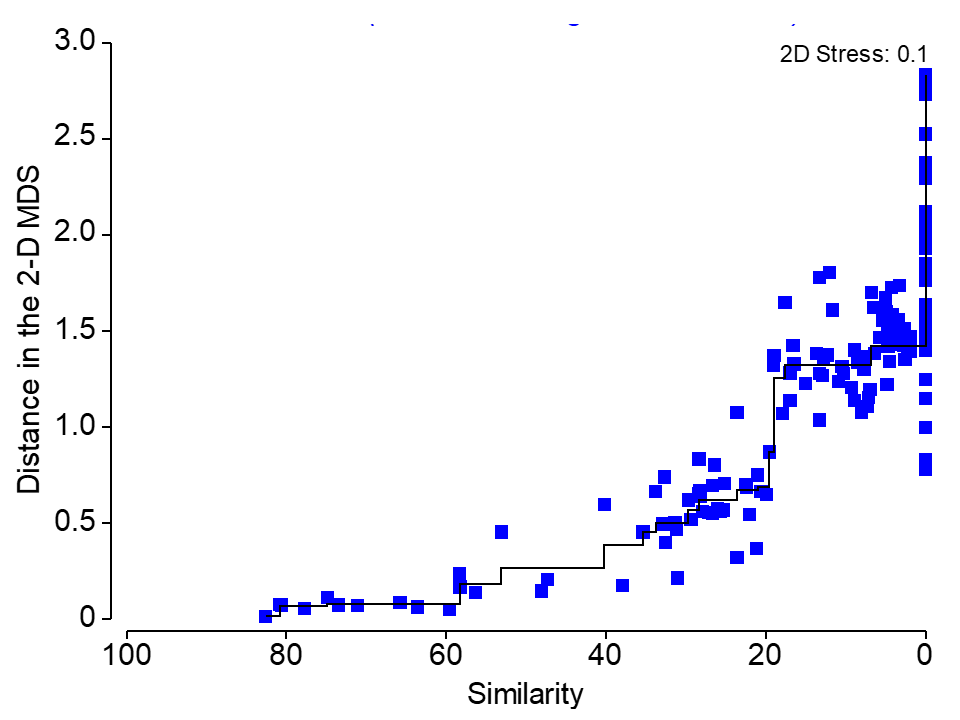
Metric VS Non-metric MDS (nMDS)
In a nMDS, points are positioned according to their ranked distances.
Therefore, the specific values along the axes of a nMDS plot have no particular meaning – they are arbitrary.
nMDS is flexible. You can use any measure of distance, dissimilarity or similarity may be used as the basis of the analysis.
nMDS is generally more robust to outliers.
nMDS tends to be more computationally intense.
Considerations about MDS
Stress will decrease with increasing numbers of MDS axes
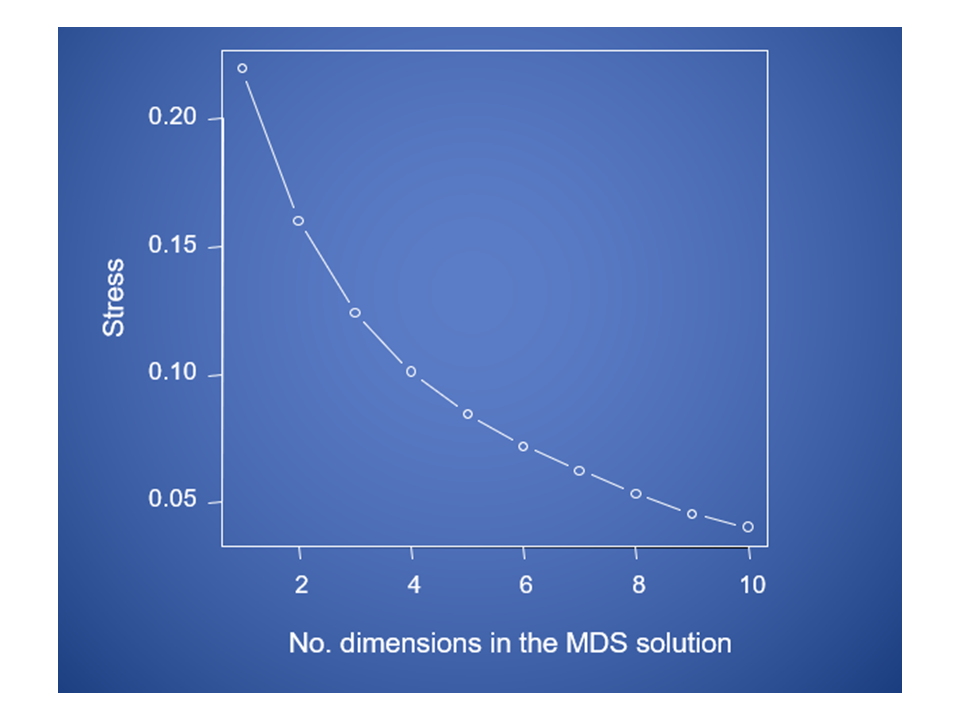
Considerations about MDS
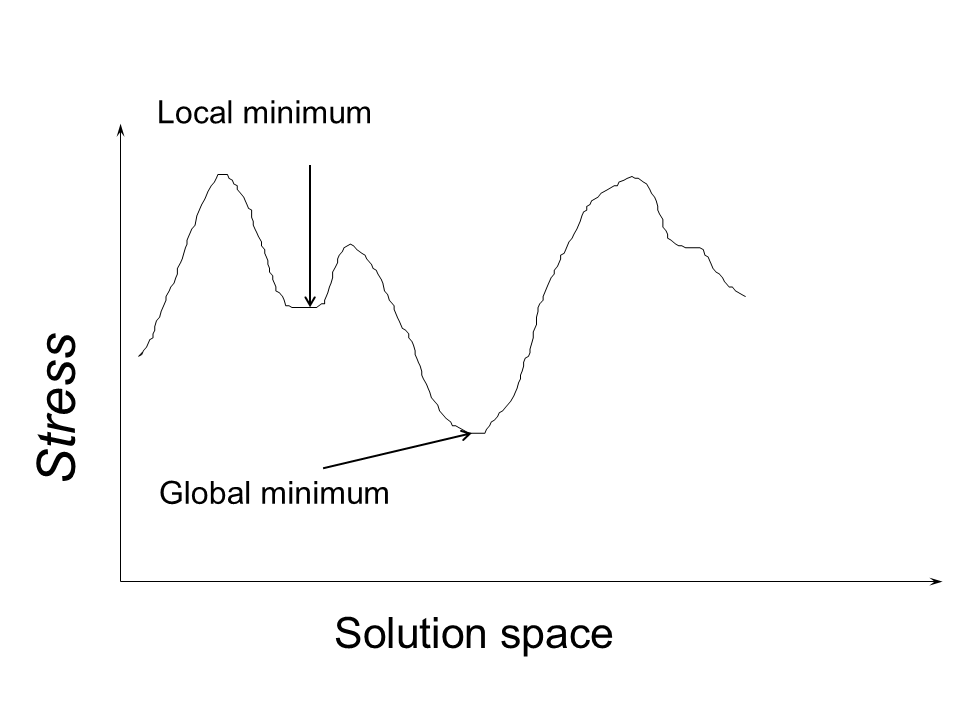
Stress can fall into local minima.
Factor Analysis (FA)
Back to the var-covar (R) matrix.
In PCA, R is eigen decomposed.
In FA, R-U. where U is the unique part. R-U is known as Communality.
\[ \begin{bmatrix} 1 & r_{12} & \cdots & r_{1p} \\ r_{21} & 1 & \cdots & r_{2p} \\ \vdots & \vdots & \ddots & \vdots \\ r_{p1} & r_{p2} & \cdots & 1 \end{bmatrix} - \begin{bmatrix} u_1 & 0 & \cdots & 0 \\ 0 & u_2 & \cdots & 0 \\ \vdots & \vdots & \ddots & \vdots \\ 0 & 0 & \cdots & u_p \end{bmatrix} \]
- FA analyzes only the common variance of the observed variables.
Factor Analysis (FA) considerations
Since the concept of common variance is hypothetical, we never know exactly in advance what percentage of variance is common and what proportion is unique among variables.
Factor scores are not linear combinations of the variables. They are estimates of latent factors. Try to avoid data fishing problems by doing the following:
- Carefully selecting your manifest variables
- Using rotation to interpret the factors
- Performing a confirmatory factor analysis to test
hypotheses about the adequacy of the factor solution
